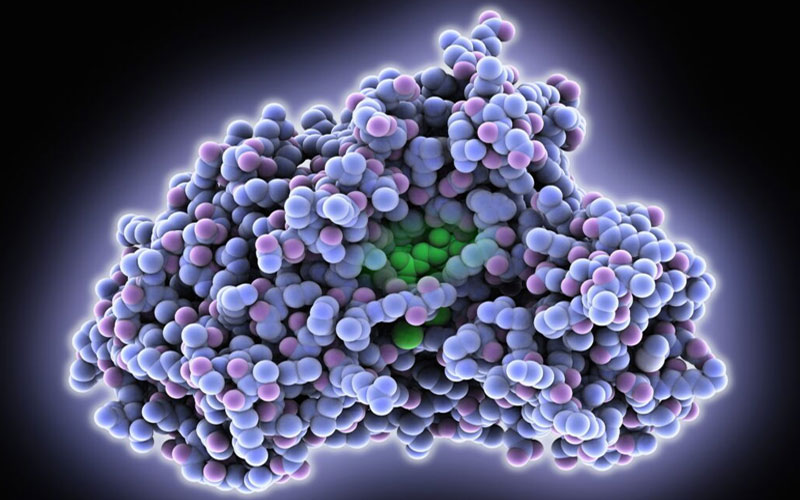
The biological and physical characteristics of Fluc-mRNA LNPs are made using various mixing techniques. The particle size distribution and cryo-EM characterization revealed multiple lipid bilayer structures. The term “FLuc mRNA” refers to Firefly Luciferase messenger RNA. Firefly Luciferase is a bioluminescent enzyme, and its mRNA is a molecule that plays a crucial role in protein synthesis.
uORFs
Upstream open reading frames (uORFs) are non-coding regions in 5′ untranslated leader sequences of mRNAs that can cause repression or enhancement of the downstream CDS depending on the context. For example, uORFs in the carnitine palmitoyltransferase 1C gene (CPT1C) 5′ leader sequence repress translation of the CPT1C protein under physiological conditions. Still, this repression is relieved by the phosphorylation of eIF2 during glucose deprivation or palmitate-BSA treatment.
A uORF’s sequence context determines whether ribosomes stall or continue scanning in the same direction, eventually recognizing and translating the downstream structural gene. Mutations and polymorphisms that disrupt or create uORFs in human genes are associated with rare diseases.
According to several studies, uORFs’ tendency to be more repressive than CDSs within the same gene may be connected to their proximity to the structural gene’s start codon. However, this is only sometimes the case, and results show that uORFs are generally under weak purifying selection in both humans and flies.
dsRNAs
Protein synthesis is a costly, energetic process that needs to be closely regulated. Thus, cells aggressively contain it by limiting the amount of RNA that gets translated. This is accomplished by triggering RNA interference (RNAi). In mammals, the most common source of dsRNA is from Short Interspersed Nuclear Elements (SINEs). These are found in all genomes and, in particular, in C. elegans, where they can be induced to trigger RNAi by feeding the little worm exogenous dsRNA.
This is a highly efficient way to induce RNAi, but it can also result in the translation of truncated protein fragments. To avoid this problem, a bioluminescent reporter gene like luciferase (Fluc) is used in most experimental settings. FLuc is a susceptible and efficient reporter gene that can be expressed directly in the cytoplasm without a promoter.
Both complementation and reconstitution strategies can be used to recover fluorescence from FLuc mRNA. In the complementation strategy, one fragment of FLuc is connected with the N-terminal of a protein splicing system, such as DNA polymerase III, and the other to the C-terminal of the same protein. The interaction between these proteins restores enzymatic activity.
mORFs
Molecular recognition features (MoRFs) are disordered regions in proteins that undergo disorder-to-order transitions upon binding to their protein partners. They are widespread in proteomes across 868 species in eukaryota, bacteria, and archaea and are commonly found in protein-protein interaction domains. Prediction of MoRFs is crucial for understanding their functional roles and identifying potential drug targets.
State-of-the-art disorder predictors have been developed to identify putative MoRFs in a sequence. The black areas in the sequence indicate regions predicted to be disordered by three popular disorder predictors. The sensitivity and specificity of these predictions are reported in the figure.
tRNAs
tRNAs are the messengers that bring amino acids to ribosomes during protein synthesis. They do this by matching the sequence of codons on mRNA to specific amino acid residues in a tRNA called an anticodon.
A tRNA can carry one or more amino acids, and its structure is shaped like a clover leaf (secondary) or an inverted L shape (tertiary). The tRNA has two RNA double helices that bend together. One has a TyC loop that pairs with some nucleotides in the mRNA codons, and another has a CCA acceptor arm.
A peptide bond is created when a tRNA’s anticodon end attaches to the matching amino acid residue in the ribosome when it aligns with an mRNA codon. tRNA molecules then leave the ribosome and move to other parts of the cell, carrying amino acids to be linked into proteins. A battery of enzymes known as aminoacyl-tRNA synthetases charge the tips of tRNA with an appropriate amino acid for the next step in protein construction.
mRNAs
When a cell wants to make a protein, it reads the DNA blueprints and directs an enzyme to copy them into a mobile intermediate message, known as messenger ribonucleic acid (mRNA). This mRNA is then transported across the cell, where it is translated into the proteins that the cell needs.
Once mRNA has been transcribed, it can be degraded or modified by protein kinases and phosphatases. These modifications are essential in controlling translation efficiency since they affect the binding of tRNA and the 40S ribosomal subunit to mRNA at the translation initiation site.
FLuc mRNA allows researchers to monitor the gene expression level in cells without the need for genetically engineered knockout mice. The mRNA transfects into the cells, translating it into luciferase protein directly in the cytoplasm. The resulting luminescence is highly sensitive and easy to measure, making it an ideal tool for longitudinal studies. It also can be used to screen for compounds that affect mRNA stability or translation.